M-ERA.NET Call 2025 – Functional materials Duration: 36 months · TRL 1 → 3
Coordinator: Medical University of Białystok, Poland
Designing programmable, curli-based amyloid materials through molecular grammar, simulations and biofabrication.
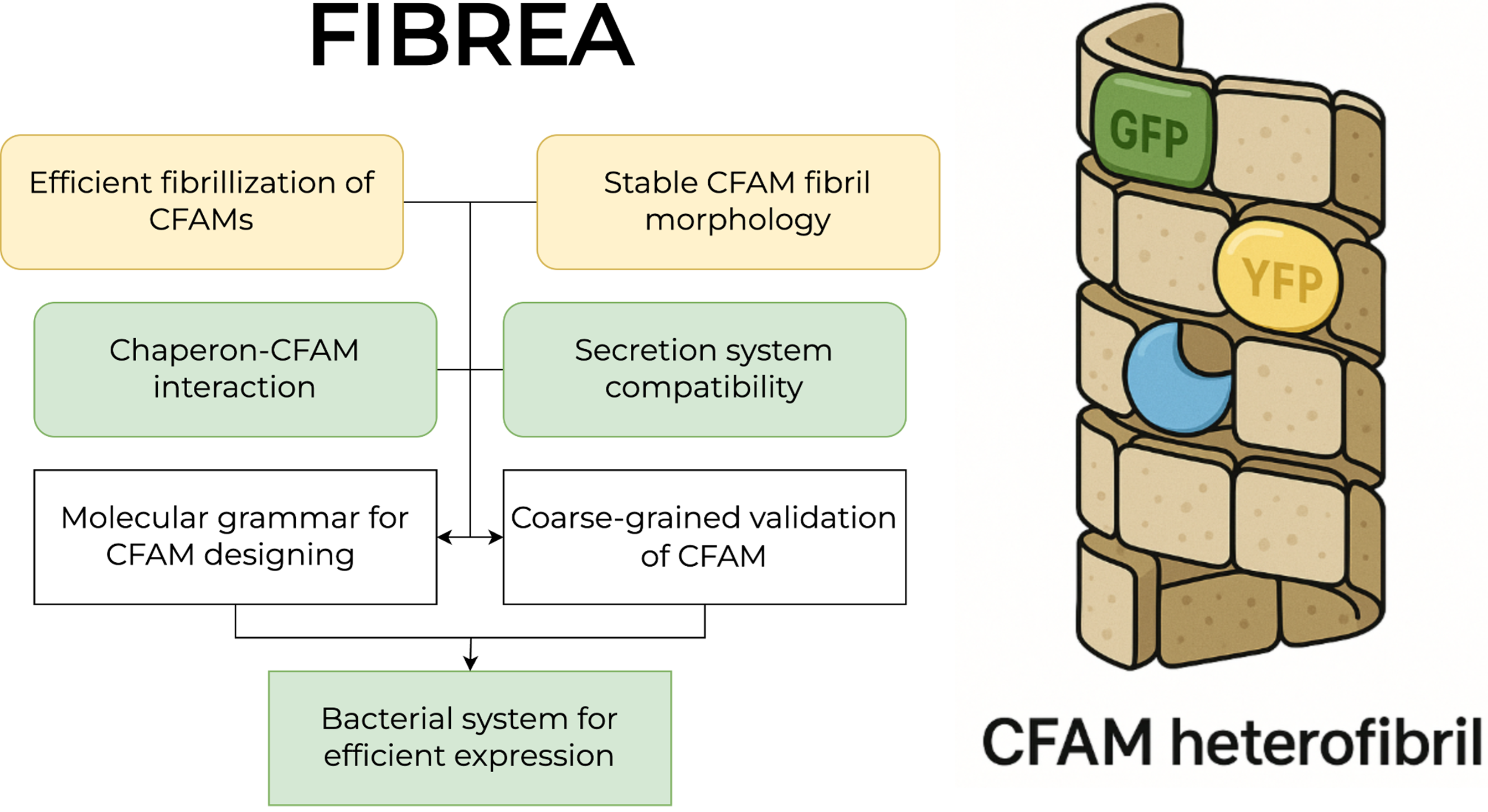
Project overview
FIBREA tackles a central challenge in functional materials: how to design amyloid-based biomaterials rationally, rather than by trial and error.
We focus on curli-based functional amyloid materials (CFAMs), engineered variants of CsgA that:
- self-assemble into fibrils with controllable morphology
- remain compatible with bacterial secretion and intracellular inhibition
- carry modular functional domains such as fluorophores or enzymes
The core idea is to learn a molecular grammar that links CFAM sequence features to fibril properties, co-assembly behavior, and production constraints.
Project at a glance
- Acronym: FIBREA
- Call: M-ERA.NET 2025 – functional materials
- Duration: 36 months
- TRL: 1 → 3
- Main outputs:
- Molecular grammar for CFAM design
- Coarse-grained simulation framework (CALVADOS-based)
- Optimized bacterial expression system
- Fluorescent and catalytic CFAM prototypes
- Molecular grammar for CFAM design
Objectives
FIBREA builds a full design–build–test–learn loop for amyloid materials.
- Develop a generalizable molecular grammar for stable CFAMs with tunable self-assembly.
- Encode compatibility with:
- curli secretion machinery (CsgEFG)
- intracellular chaperones (CsgC, Spy)
- Model CFAM assembly and inhibition using coarse-grained simulations (CALVADOS).
- Optimize a bacterial expression system for high-yield CFAM production.
- Demonstrate:
- fluorescent CFAMs (GFP/YFP, including FRET-based co-assembly)
- enzymatic CFAMs for multi-step catalytic cascades.
- Advance the overall system from TRL 1 to TRL 3.
Concept and approach
FIBREA combines AI, physics-based simulations and wet-lab validation into a single workflow.
- Molecular grammar (WP1)
- Conditional denoising diffusion model trained on > 43 000 curli operons.
- Encodes secretion compatibility, inhibition susceptibility, and fibrillization behavior.
- Conditional denoising diffusion model trained on > 43 000 curli operons.
- Coarse-grained simulations (WP2)
- CALVADOS used to simulate nucleation, growth, and polymorphism of CFAM fibrils.
- Outputs structural descriptors (contact maps, stiffness, solvent exposure) feeding back into the grammar.
- CALVADOS used to simulate nucleation, growth, and polymorphism of CFAM fibrils.
- Expression and production (WP3)
- Artificial gene design, codon optimization, fermentation strategy comparison, purification workflows.
- Structural and biophysical validation (WP4)
- ThT, Congo Red, Amytracker, CD, FTIR, TEM, AFM, cryo-EM.
- In vitro testing of CsgC variants as inhibitors.
- ThT, Congo Red, Amytracker, CD, FTIR, TEM, AFM, cryo-EM.
- Functional CFAMs (WP5)
- Fluorescent CFAMs for tracking and FRET analyses.
- Dual-enzyme CFAMs as molecular assembly lines.
- Fluorescent CFAMs for tracking and FRET analyses.
Only independently reproduced fibrillization results (Lithuanian labs) are used to refine the grammar, enforcing robustness.
Impact
Scientific
- Establishes design rules linking sequence → structure → function in amyloid materials.
- Bridges machine learning, coarse-grained physics, and experiments.
- Provides new insight into the curli system and CsgC-mediated inhibition.
Economic
- Lays the foundation for industrial CFAM applications: biosensing, catalysis, filtration, remediation.
- Enables bio-based, programmable materials as alternatives to petroleum-derived polymers.
Societal and environmental
- Promotes biodegradable, low-toxicity materials from microbial systems.
- Implements FAIR data, Open Science, and Responsible Research and Innovation (RRI).
- Engages the public to show amyloids beyond their disease associations.
Consortium
FIBREA unites complementary expertise across bioinformatics, simulations, amyloid biology and synthetic biology.
Leads molecular grammar development and overall coordination. Expertise in protein informatics, machine learning for amyloids and peptides and FAIR/DOME-compliant data workflows.
Leader:
Experimental amyloid specialists performing biophysical characterization, structural validation and independent reproducibility checks.
Leader:
Developers of CALVADOS and world leaders in protein folding and aggregation modeling. Provide physics-based constraints and structural descriptors for CFAMs.
Leader:
Experts in curli-based nanomaterials and biofilm engineering. Optimize host strains, codon usage, fermentation and purification for CFAM production.
Leader:
Focus on functional CFAMs carrying fluorescent and enzymatic modules. Demonstrate catalytic CFAMs and FRET-based co-assembly.
Leader:
Work packages
The work plan is organized into six interlinked WPs, forming a closed design–build–test–learn loop.
- Define biological constraints for secretion-competent constructs.
- Learn CsgA variability and co-evolution with CsgC.
- Develop conditional diffusion models for CFAM design.
- Integrate feedback from simulations (WP2) and experiments (WP3–5).
- Set up CALVADOS simulations for CFAMs and inhibitors.
- Screen engineered sequences in silico.
- Extract structural descriptors for morphology and stability.
- Calibrate against data from WP3 and WP4.
- Select host systems and design synthetic genes.
- Optimize codon usage and expression constructs.
- Compare batch vs continuous fermentation.
- Improve purification and confirm functional integrity.
- Biophysical assays (ThT, Congo Red, Amytracker, CD, FTIR).
- TEM and AFM imaging of fibrils.
- Cryo-EM structures of selected CFAM fibrils.
- Inhibition studies with CsgC variants.
- Assess amyloid aggregation of functional CFAM fusions.
- Validate GFP/YFP-based co-assembly and FRET.
- Demonstrate dual-enzyme CFAM catalytic cascades.
- Test inter-lab reproducibility of fibrillization and activity.
- Scientific, administrative, and financial coordination.
- Mattermost-based internal communication and data sharing.
- Open Science, FAIR data, and RRI implementation.
- Website, outreach, and training for early-career researchers.
Sustainability, RRI and data management
- Sustainability:
- Protein-based, biodegradable materials; microbial expression; potential for circular use.
- Focus on low-energy, resource-efficient production strategies.
- Protein-based, biodegradable materials; microbial expression; potential for circular use.
- RRI and ethics:
- Ethical review of experimental activities where required.
- Biosecurity risk assessment aligned with EU and national regulations.
- Public engagement on the constructive uses of amyloids.
- Ethical review of experimental activities where required.
- Data and open science:
- FAIR-compliant data, with deposition to public repositories (e.g. Zenodo, NCBI).
- Code release (e.g. GitHub) under permissive licenses.
- Documentation following the DOME recommendations for machine learning in life sciences.
- FAIR-compliant data, with deposition to public repositories (e.g. Zenodo, NCBI).
Publications and outputs
A detailed list will be maintained as the project progresses. Planned outputs include:
- Articles on:
- CFAM molecular grammar and generative modeling
- CALVADOS-based CFAM simulations
- Structural and functional characterization of CFAMs
- CFAM molecular grammar and generative modeling
- Conference presentations and workshops.
- Open-source tools and curated datasets.
Contact
Project Coordinator
Michał Burdukiewicz, PhD
Medical University of Białystok
Jana Kilińskiego 1, 15-089 Białystok, Poland