Short amyloids ≠ antimicrobial peptides? 🧬❌🦠
amyloids, antimicrobial peptides, AMP, machine learning, AmpGram, negative dataset, bioinformatics
📌 Project highlights
- 🧬 Tests short amyloids predicted as AMPs
- 🤖 Uses AmpGram + 15 ML models
- 🧪 Validates experimentally on 10 bacterial strains
- ❌ Finds no antimicrobial activity
- 📊 Proposes high-quality negative dataset for ML
🎉 New paper!
👉 testing if amyloids can act like antimicrobial peptides
👉 Testing Antimicrobial Properties of Selected Short Amyloids
🎧 Audio summary
Machine learning said “these might be AMPs”…
experiments said “nope” 😅
👉 Here’s a quick breakdown 🎧:
🔬 What is this about?
Amyloids and antimicrobial peptides (AMPs) look surprisingly similar:
- both can disrupt membranes
- both form aggregates (β-sheet structures)
- both interact with the immune system
👉 So the question: Can short amyloids act as antimicrobial peptides?
⚙️ The core problem
AMP discovery relies heavily on machine learning
BUT:
👉 models lack true negative examples
- very few experimentally confirmed non-AMPs
- datasets often contain false negatives
👉 leads to over-optimistic predictions
🧠 What they did
🧩 Step 1: ML-based selection
- screened 509 amyloids (WALTZ-DB)
- used AmpGram (n-grams + random forest)
- selected top 10 candidates
👉 only ~6% predicted as AMPs
⚖️ Step 2: model comparison
- tested 15 additional AMP predictors
👉 result:
- highly inconsistent predictions
- only 1 peptide agreed across all models
🧪 Step 3: experimental validation
They tested:
- 🦠 10 bacterial strains (Gram+ & Gram−)
- 🧫 peptide concentrations up to 128 µg/mL
- 🧬 human cell toxicity
🔍 Key results
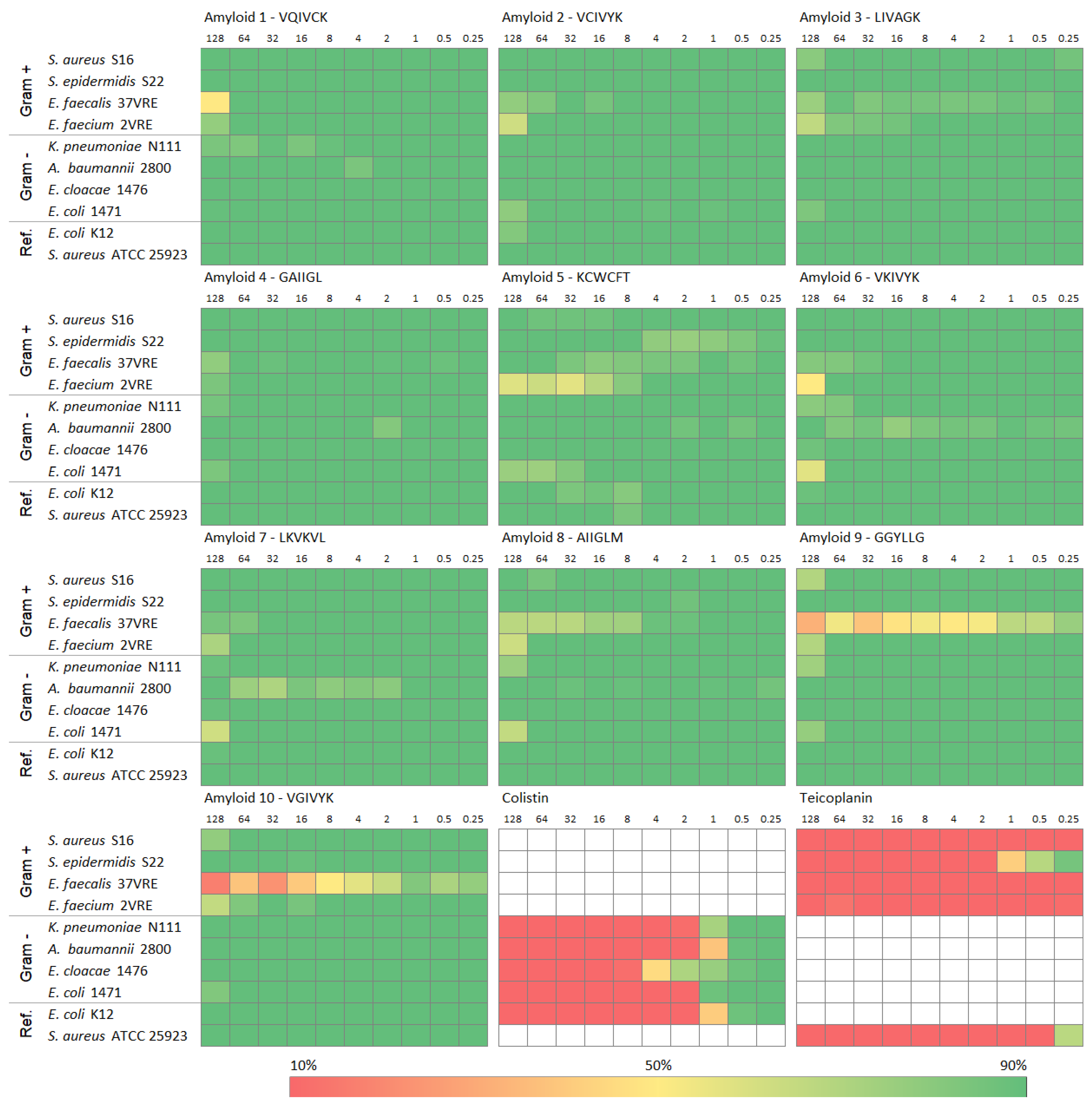
❌ No antimicrobial activity
- all peptides remained green (bacteria survived)
- MIC > 128 µg/mL → no effective killing
👉 even top ML predictions failed
🧬 No cytotoxicity either
- no harmful effects on human cells
- IC50 values relatively high
👉 peptides are biologically inactive (safe but useless as AMPs)
🧩 Some amyloids still aggregate
- 4/10 formed fibrils
- aggregation ≠ antimicrobial activity
👉 structural similarity ≠ functional similarity
⚠️ ML models struggle here
- short amyloids = edge cases
- predictions vary wildly across tools
👉 shows limits of current AMP predictors
💡 Key insight
👉 This is a negative result and that’s the point
These peptides:
- look like AMPs
- are predicted like AMPs
- behave like AMPs structurally
BUT:
👉 they are NOT AMPs
🚀 Why this matters
🧠 Better ML models
These sequences are:
👉 perfect “hard negatives”
- highly similar to AMPs
- experimentally validated as non-AMPs
👉 ideal for training robust models
⚠️ Dataset problem
Only ~24 confirmed non-AMPs exist (!!!)
👉 massive bottleneck in AMP prediction
📣 Cultural shift in science
The paper strongly argues:
👉 publish negative results
- prevents bias
- improves ML datasets
- accelerates discovery
💚 BioGenies perspective
This paper is 🔥 for one reason:
👉 it exposes a hidden failure mode in bio-ML
- models ≠ biology
- predictions ≠ function
- similarity ≠ activity
👉 And more importantly:
negative data = high-value data
This is exactly what:
- 🧠 interpretable ML
- 🧬 robust datasets
- 🔬 real-world validation
should look like