aSynPEP-DB: mining peptides against Parkinson’s 🧬🧠
alpha-synuclein, Parkinson’s disease, peptides, database, bioinformatics, AMP, neuropeptides
📌 Project highlights
- 🧠 Targets α-synuclein aggregation (Parkinson’s hallmark)
- 🧬 Screens neuropeptides, AMPs, microbiome + food peptides
- 🤖 Uses physicochemical rules-based algorithm
- 📊 Identifies 123 candidate inhibitory peptides
- 🌐 Provides interactive database + prediction tool
🎉 New paper!
👉 building a peptide database to fight Parkinson’s
👉 aSynPEP-DB: a database of biogenic peptides for inhibiting α-synuclein aggregation
🔗 Try it yourself
🎧 Audio summary
What if your body already contains molecules
that can slow Parkinson’s disease? 🤯
👉 This paper builds a database to find them 🎧
🔬 What is this about?
Parkinson’s disease (PD) is driven by:
👉 aggregation of α-synuclein (aSyn) into toxic species
BUT:
- no therapies stop aggregation
- designing molecules is hard
- aSyn is intrinsically disordered
👉 difficult drug target
💡 Key idea:
Some natural peptides can:
- bind toxic aSyn oligomers
- block aggregation
- reduce toxicity
👉 so… can we systematically find them?
⚙️ The core concept
The study builds: 👉 aSynPEP-DB
A database of peptides predicted to: 👉 inhibit α-synuclein aggregation
📊 It integrates:
- 🧠 human neuropeptides
- 🦠 antimicrobial peptides (AMPs)
- 🧫 gut microbiome peptides
- 🥛 food-derived bioactive peptides
🧠 The key insight
From prior experiments (e.g. LL-37):
👉 active peptides share 3 properties:
- 🌀 α-helical structure
- ⚖️ amphipathicity
- ➕ positive net charge
These allow binding to 👉 negatively charged, hydrophobic aSyn aggregates
🧩 What they did
🔍 Step 1: dataset collection
From multiple databases:
- NeuroPep (neuropeptides)
- DRAMP (AMPs)
- GMrepo (microbiome)
- DFBP (food peptides)
👉 thousands of peptides screened
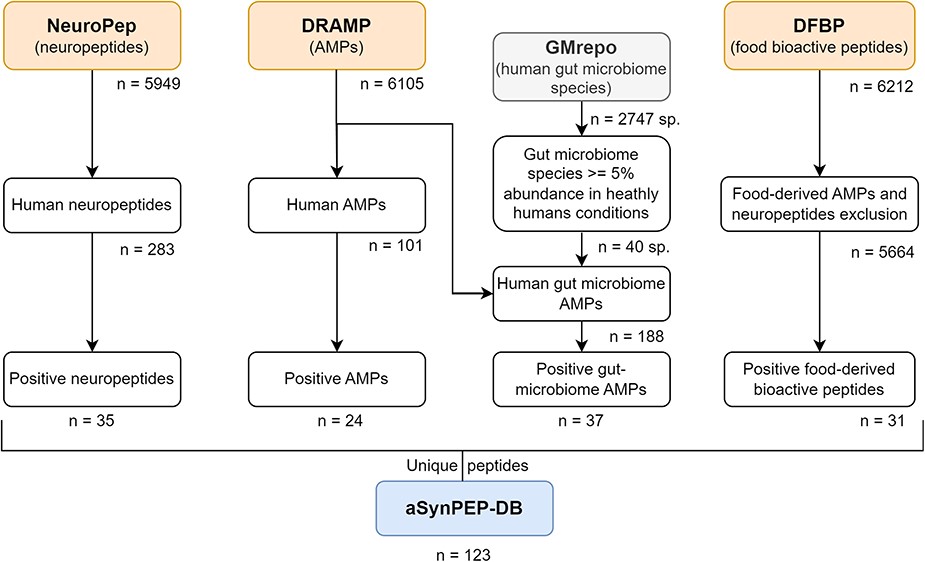
🤖 Step 2: discriminative algorithm
Heuristic filtering based on:
- α-helical propensity (AGADIR)
- amphipathicity (hydrophobic moment)
- net charge
👉 also scans sub-sequences (sliding window)
🧬 Step 3: candidate selection
👉 123 unique peptides identified
🌐 Step 4: database construction
Each entry includes:
- sequence + inhibitory region
- structure (AlphaFold)
- toxicity prediction
- BBB permeability
- tissue expression
🔍 Key results
🧬 A new peptide landscape
- 123 candidate inhibitors
- spanning multiple biological sources
👉 many previously unexplored
🧠 Biologically relevant hits
Examples include:
- Neuropeptide Y (NPY) → neuroprotective, brain-expressed
- BMAP-28 → antimicrobial + food-derived
- Lactoferricin-H → immune-related peptide
- OR-7 (microbiome) → gut-brain axis relevance
⚠️ Most are NOT experimentally validated
👉 database = hypothesis generator
- only a few peptides (e.g. LL-37) validated
- majority remain predictions
💡 Key insight
👉 Nature already encodes peptides with:
- antimicrobial activity
- anti-amyloid potential
- immune modulation
👉 these functions may be evolutionarily linked
🚀 Why this matters
🧠 New therapeutic strategy
Instead of small molecules:
👉 use peptides to block aggregation
- potentially safer
- biologically compatible
- target-specific
🌍 Systems-level view
Combines:
- brain peptides
- gut microbiome
- diet
👉 connects gut–brain axis + PD
🤖 Tool for discovery
The database includes:
👉 a screening algorithm
- test new peptides
- design synthetic ones
- expand datasets
💚 BioGenies perspective
This paper is powerful because it:
👉 shifts from prediction → infrastructure
Instead of one model:
- builds a resource
- encodes biophysical rules
- enables future discoveries

