Are AMP models lying to us? 🤖🧬 The benchmarking problem
AMP, machine learning, benchmarking, negative dataset, bias, bioinformatics, reproducibility
📌 Highlights
- 🤖 Built 660 ML models across 12 architectures
- 🧪 Tested 11 negative sampling strategies
- ⚠️ Shows benchmarking in AMP prediction is biased
- 📊 Performance depends on dataset similarity (not model quality)
- 🌐 Introduces AMPBenchmark for fair evaluation
🎉 New paper out! This one tackles a very uncomfortable truth:
👉 your “state-of-the-art” AMP model might just be lucky 😄
👉 Benchmarks in antimicrobial peptide prediction are biased due to the selection of negative data
🔗 Try it yourself
👉 finally, a way to benchmark AMP models fairly
🎧 Audio summary
Many AMP predictors claim:
👉 “we outperform previous methods”
But… what if the benchmark itself is broken? 😅
👉 Here’s a short audio overview 🎧 explaining the problem:
🔬 What is this about?
Antimicrobial peptides (AMPs):
- 🧬 short bioactive peptides
- 🦠 kill bacteria, viruses, cancer cells
- 💊 promising alternative to antibiotics
👉 Because of that:
- dozens of ML predictors exist
- each claiming better performance
💡 BUT:
👉 all of them depend on training data
especially:
👉 negative data (non-AMPs)
⚠️ The core problem
There are:
- thousands of known AMPs ✅
- almost no confirmed non-AMPs ❌
👉 so researchers:
- generate artificial negatives
- using different sampling strategies
💥 Problem:
👉 these strategies create very different datasets
And ML models:
👉 learn dataset artifacts, not biology
🧠 What they did
🧪 Massive benchmark
- 12 ML architectures
- 11 negative sampling methods
- 660 models total
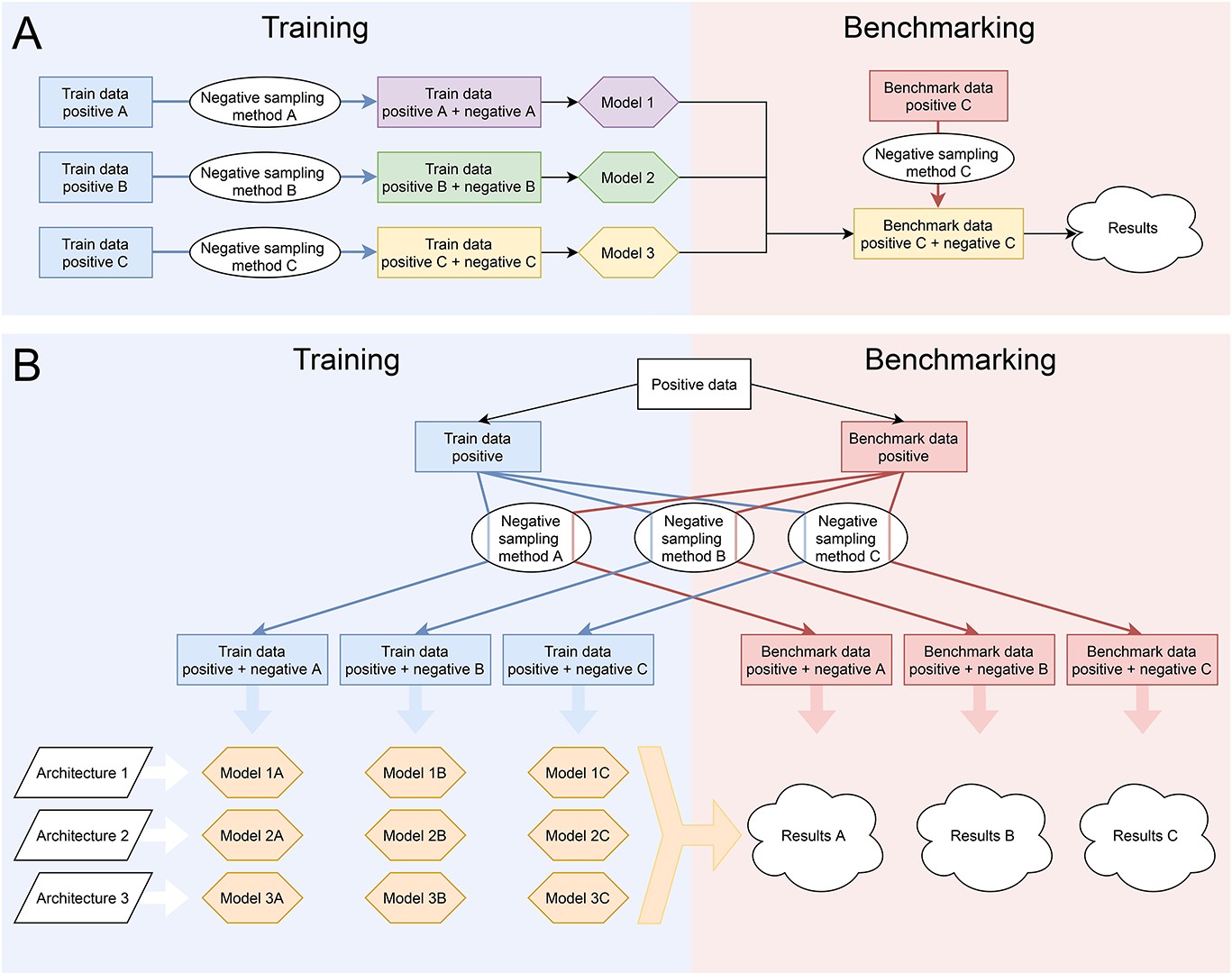
🔁 Full cross-evaluation
Each model:
- trained on one dataset
- tested on ALL others
👉 not just “friendly benchmarks”
📊 Evaluation metric
- ROC curves
- AUC scores
🔍 Key results
⚠️ Benchmarking is biased
👉 Models perform best when:
- training set = benchmark set
📉 Performance drops when:
- datasets differ
👉 meaning:
models don’t generalize, they memorize dataset structure
🧬 Dataset similarity drives performance
Strong correlations found between:
- amino acid composition similarity
- sequence length similarity
and model performance
👉 not biology
👉 just dataset artifacts
🤖 Architecture matters… but less than you think
- Random Forest models performed best
- Deep learning ≠ automatically better
👉 but dataset choice still dominates
🔁 Reproducibility crisis 🚨
- ~70% models not reproducible
- lack of code / data sharing
👉 slows down the field
💡 Key insight
👉 We don’t actually know:
which AMP model is the best
Because:
- benchmarks are biased
- comparisons are unfair
- datasets are inconsistent
👉 “state-of-the-art” = often dataset-specific
🚀 What we have built
🌐 AMPBenchmark
A platform for:
- fair model comparison
- standardized datasets
- reproducible evaluation
👉 similar idea to:
- Kaggle-style benchmarking
👉 solves:
- hidden bias
- unfair comparisons
- reproducibility issues
🚀 Why this matters
🧠 For ML in biology
This is not just AMP problem:
👉 affects:
- protein function prediction
- interaction prediction
- genomics ML
⚠️ For researchers
You should:
- question benchmarks
- test cross-dataset generalization
- avoid overfitting to dataset design
💊 For drug discovery
Bad models =
👉 missed therapeutic candidates
👉 false positives
💚 BioGenies perspective
This paper is 🔥 because it says:
👉 “your model is not as good as you think”
And more importantly:
👉 shows WHY