Amyloid cross-interactions: how proteins influence each other’s aggregation 🧬🧊
amyloids, protein aggregation, cross-seeding, cross-interactions, protein misfolding, neurodegeneration, experimental methods, bioinformatics
📌 Project highlights
- 🧬 Comprehensive review of amyloid cross-interactions
- ⚙️ Covers key experimental techniques (fluorescence, microscopy, spectroscopy)
- 🔍 Explains cross-seeding and fibril polymorphism
- ⚠️ Shows why single-method studies are insufficient
- 🧠 Recommends multi-technique strategies for reliable results
🎉 New paper out! This time, we dive into something fundamental (and tricky):
👉 how amyloids influence each other’s aggregation 😄
👉 Experimental methods for studying amyloid cross‐interactions
🎧 Audio summary
Amyloid aggregation + cross-seeding + 10 experimental techniques?
Yeah… this one can get dense 😄
👉 So we added a short audio walkthrough 🎧 to make it easier.
🔬 What is this about?
Amyloids are misfolded protein aggregates associated with many diseases:
- Alzheimer’s 🧠
- Parkinson’s ⚡
- systemic amyloidoses
But here’s the twist: 👉 amyloids don’t act alone
They can interact with each other, influencing aggregation in complex ways - a process known as:
🧩 Amyloid cross-interactions
These interactions can:
- accelerate aggregation
- inhibit fibril formation
- create new fibril structures (polymorphs)
🚨 Why this matters
Cross-interactions are central to:
- 🧠 prion-like propagation
- 🧬 co-morbidity between diseases
- ⚡ unexpected aggregation pathways
👉 In short: one protein can change the fate of another
🧠 The core problem
Studying these interactions is extremely difficult because:
- aggregation is dynamic
- structures are heterogeneous
- multiple species coexist (monomers, oligomers, fibrils)
👉 And most importantly: ❌ no single experimental method can capture the full picture
⚙️ Experimental toolbox
The paper provides a complete overview of methods used to study amyloid cross-interactions:
🌟 Fluorescence-based assays
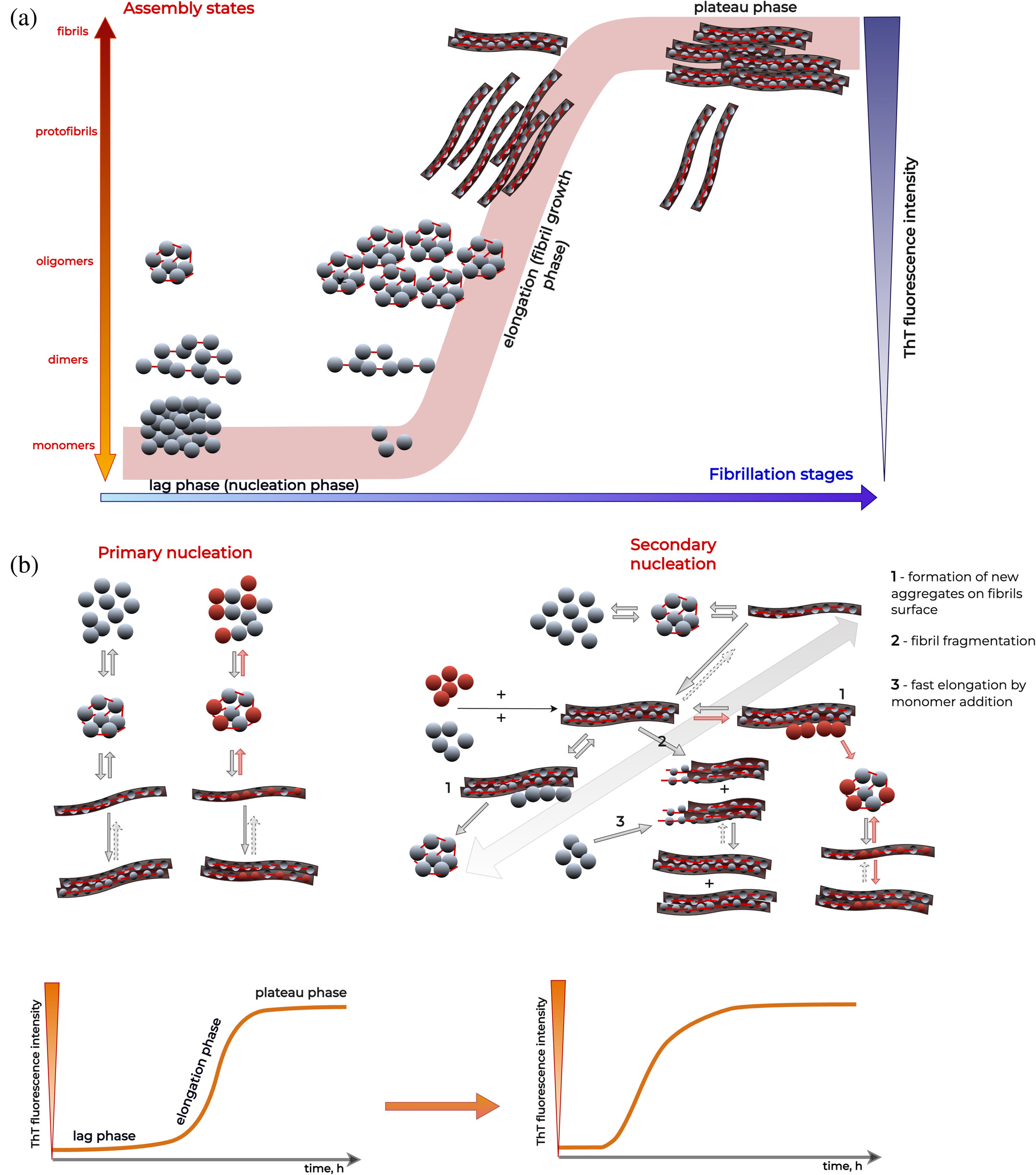
- Thioflavin T (ThT)
- Congo Red
👉 Great for:
- tracking aggregation kinetics
👉 Limitation:
- indirect, can be misleading alone
🔬 High-resolution imaging
- Atomic Force Microscopy (AFM)
- Cryo-EM
👉 Reveals:
- fibril morphology
- structural differences
🧪 Spectroscopy & structural methods
- Solid-state NMR
- Mass spectrometry
👉 Provides:
- molecular-level insights
- composition of heterotypic fibrils
🧫 Immuno-based techniques
- Immuno-EM
👉 Confirms:
- presence of mixed (cross-seeded) fibrils
🔗 The key takeaway
👉 You need a combination of methods
A typical robust workflow includes:
- fluorescence → detect aggregation
- microscopy → observe morphology
- structural methods → confirm composition
👉 Only this hybrid approach can capture the full complexity
🧬 Key biological insights
- 🧩 Cross-seeding is context-dependent
- ⚡ Same proteins can:
- accelerate
- inhibit
- or reshape aggregation
- accelerate
- 🧠 Amyloid fibrils are structurally polymorphic
- 🔄 Interactions can generate new aggregate species
👉 Translation: aggregation is not a single pathway - it’s a network
🚀 Why this matters (big picture)
This review helps:
- 🧠 design better experiments
- 🔬 interpret conflicting results
- ⚙️ improve models of aggregation
- 💊 guide therapeutic strategies
👉 Especially important for:
- multi-protein diseases
- cross-talk between aggregation pathways
💚 BioGenies perspective
This is exactly in our playground:
- amyloids 🧊
- aggregation prediction 🧠
- phase separation ⚡
👉 And reinforces a key idea we keep seeing you cannot understand aggregation without considering interactions