LLPS datasets & benchmarking: bringing order to condensate chaos 🧬📊
LLPS, liquid-liquid phase separation, biomolecular condensates, protein datasets, machine learning, bioinformatics, benchmarking
📌 Project highlights
- 🧬 First integrated datasets of LLPS proteins (drivers, clients, negatives)
- ⚖️ Introduces standardized negative datasets (structured + disordered proteins)
- 📊 Benchmark of 16 LLPS prediction tools - the most comprehensive to date
- 🧠 Reveals limitations of current ML models for phase separation
- 🌐 Public resource: datasets + website for reproducible research
🎉 New paper out! And this one is a big one, not just another tool, but a foundation for the whole field 😄
👉 Comprehensive protein datasets and benchmarking for liquid–liquid phase separation studies
🔗 Data & resources
This is not just a paper, it’s a community resource for building and benchmarking LLPS models.
🎧 Audio summary
Not everyone wants to read about κ parameters and sticker–spacer distributions over coffee ☕😄
👉 We’ve attached a short audio explanation 🎧 to make things easier.
🔬 What is this about?
Liquid–liquid phase separation (LLPS) is a process where proteins form biomolecular condensates, membrane-less compartments that organize cellular processes.
These condensates are crucial for:
- gene regulation
- stress response
- disease mechanisms (e.g. neurodegeneration)
But studying LLPS computationally has a major problem:
👉 data is messy, inconsistent, and fragmented across databases
🚨 The problem we tackled
Existing LLPS databases:
- use different definitions
- contain inconsistent annotations
- lack reliable negative examples
👉 This makes:
- machine learning unreliable
- benchmarking unfair
- results hard to compare
🧠 What we built
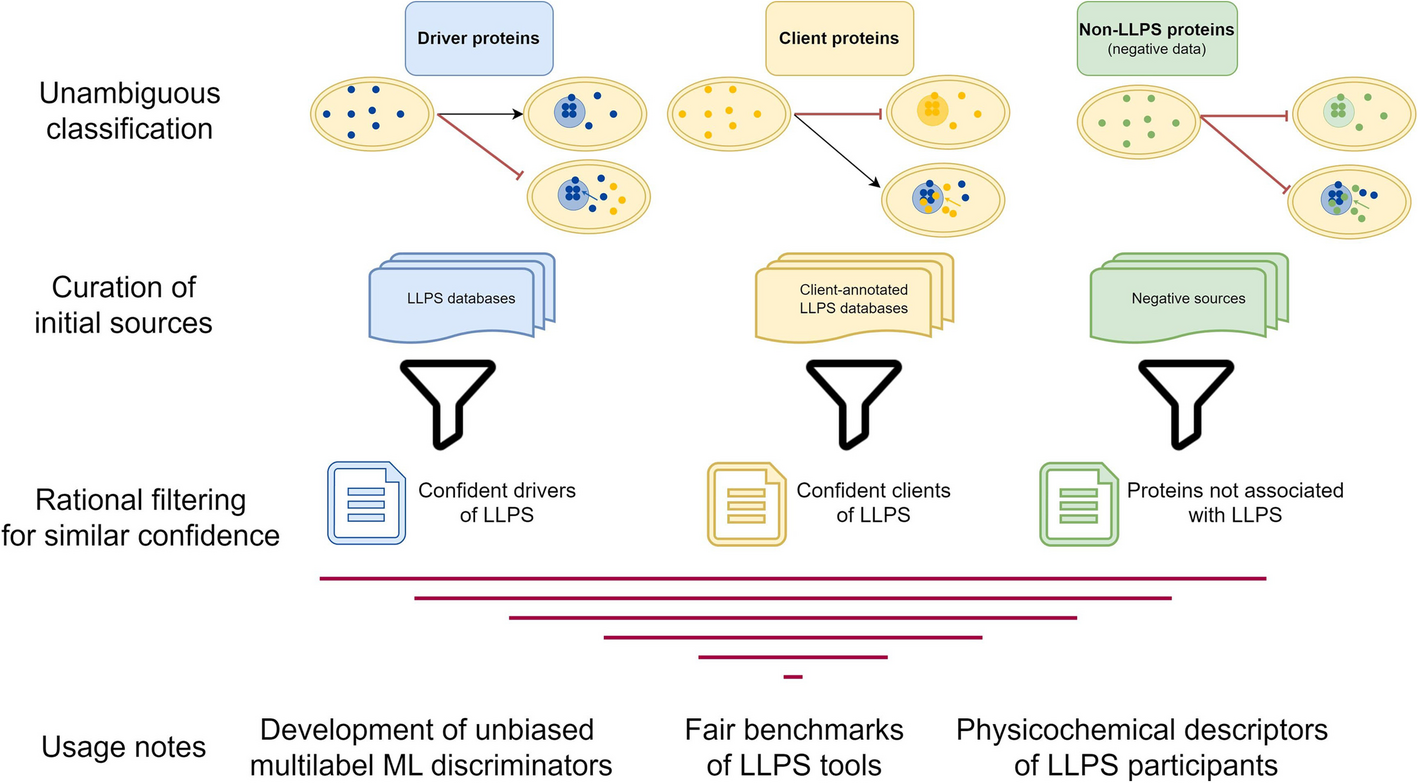
We created high-confidence datasets of proteins involved in LLPS, based on:
🧬 Protein roles
- Drivers → form condensates independently
- Clients → join existing condensates
- Negatives → proteins not associated with LLPS
👉 Importantly, we introduced two types of negative datasets:
- structured proteins (PDB-like)
- disordered proteins (DisProt-like)
This is critical because: 👉 disorder alone ≠ LLPS (big source of bias!)
⚙️ How it works
We used:
- 📚 integration of multiple LLPS databases
- 🧹 strict filtering criteria for data quality
- 🧬 annotation of:
- disorder
- prion-like domains
- physicochemical features
- disorder
Then we:
👉 analyzed protein properties
👉 and benchmarked prediction tools
📊 Benchmarking LLPS predictors
We evaluated 16 bioinformatics tools on independent datasets.
Key findings:
- 🤖 ML-based tools perform best
- ⚠️ Many tools still have high false positive rates
- 🧠 Models often confuse:
- disorder
- with actual phase separation
- disorder
👉 Translation: we are still not great at predicting LLPS reliably 😄
🧬 Key biological insights
From the analysis:
- 🧩 LLPS is multi-factorial, not driven by a single feature
- ⚡ properties like:
- charge distribution
- aggregation propensity
- interaction regions
matter more than simple sequence features
- charge distribution
- 🧠 drivers and clients are different, but subtle
👉 And importantly: you cannot model LLPS properly without good negative data
🚀 Why this matters
This work:
- 📊 sets a new standard for benchmarking
- 🧠 improves machine learning reliability
- 🔬 enables better biological interpretation of condensates
👉 In short: we finally have a clean dataset to stop guessing and start learning properly
💚 BioGenies context
This project connects directly to our interests in:
- amyloids 🧬
- protein aggregation 🧊
- phase separation ⚡
👉 and gives us a solid foundation for future tools and models
💚 BioGenies perspective
This project connects directly to our interests in:
- amyloids 🧬
- protein aggregation 🧊
- phase separation ⚡
👉 and gives us a solid foundation for future tools and models