ML + metabolomics in the clinic: promise vs reality 🧠🧬
metabolomics, machine learning, clinical decision support, CDSS, biomarkers, precision medicine, AI
📌 Project highlights
- 🧠 Machine learning for clinical decision support systems (CDSS)
- 🧬 Focus on metabolomics data complexity
- ⚙️ Covers full pipeline: data → models → clinical utility
- ⚠️ Critical discussion of limitations & risks
- 🚀 Outlook on precision medicine applications
🎉 New review out! This one sits at the intersection of AI, metabolomics and clinical practice
🔗 Explore the paper
👉 A deep dive into how machine learning turns metabolomics into clinical decisions.
🎧 Audio summary
Not everyone wants to dive straight into ML, metabolomics, and clinical pipelines (fair 😄)
👉 Here’s a short audio walkthrough 🎧 explaining what this work is about and why it matters:
🔬 What is this about?
Modern medicine generates massive, multi-layered datasets.
Among them:
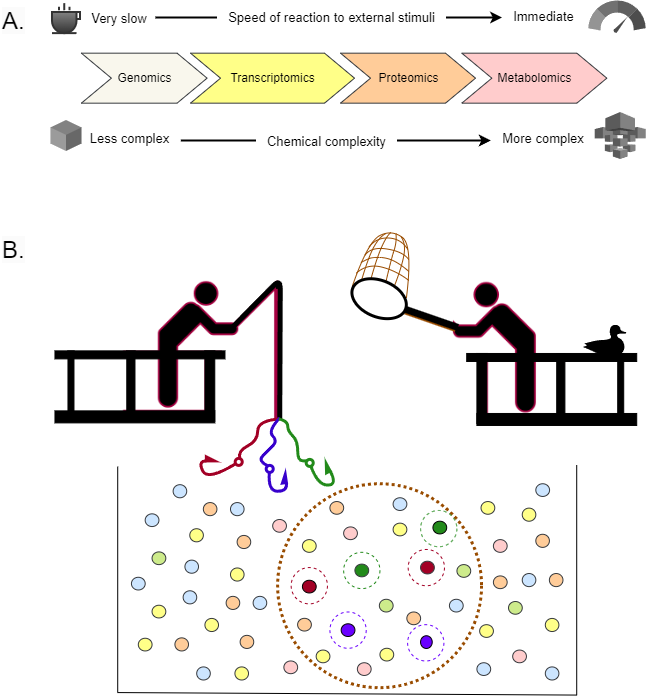
👉 metabolomics captures the current physiological state of a patient
- fast response to environmental changes
- extremely high chemical diversity
- thousands of measurable molecules
But this comes at a cost:
❗ extreme complexity → requires machine learning
🧠 Enter: Clinical Decision Support Systems (CDSS)
ML-based CDSS aim to:
- diagnose diseases
- predict outcomes
- guide treatment decisions
👉 essentially simulate clinical reasoning using data
Typical pipeline:
- 🧪 Sample collection (blood, urine, tissue)
- 🔬 MS / NMR → metabolite profiles
- 📊 Data processing
- 🤖 ML model → prediction
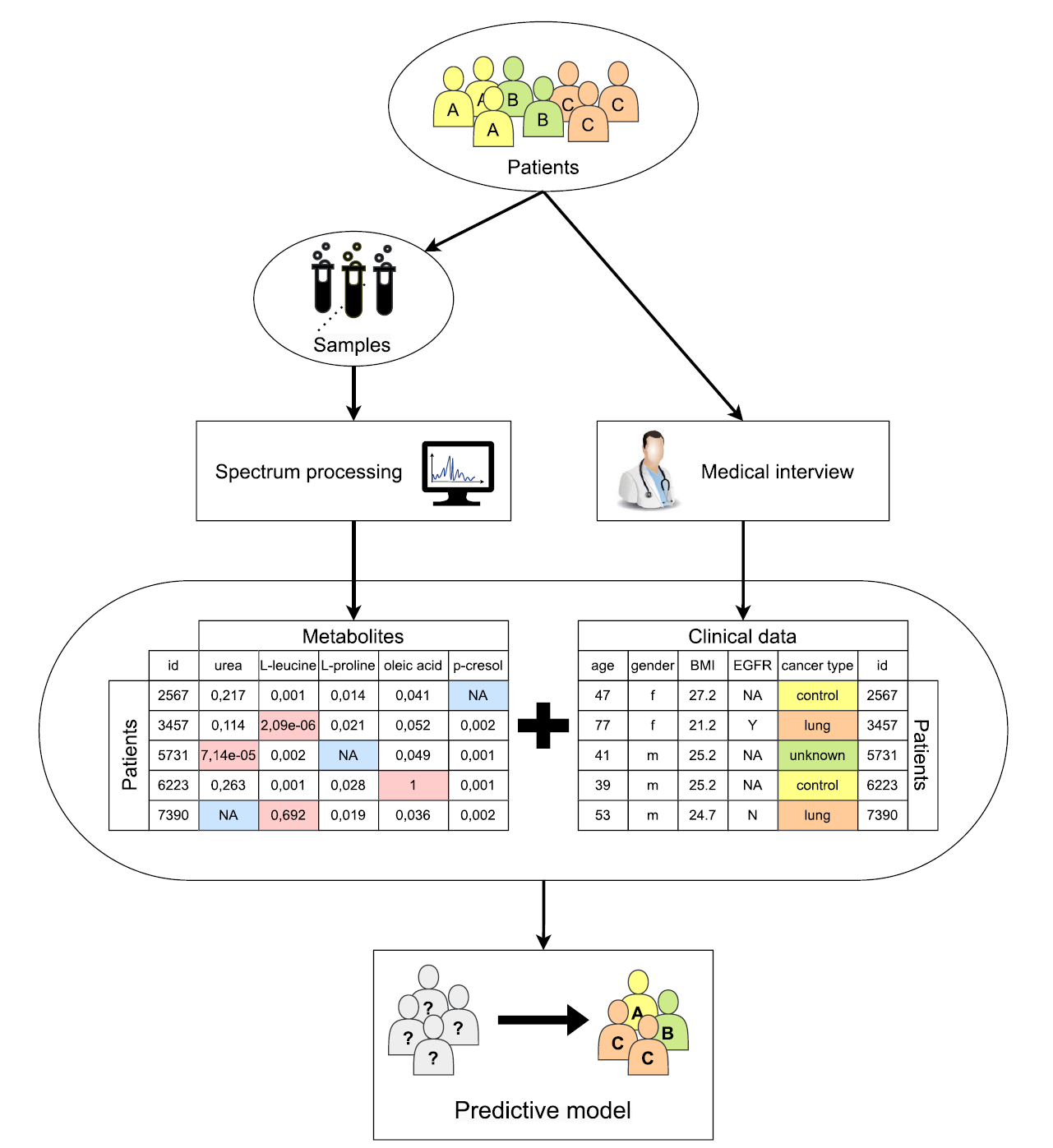
📊 The diagram shows this full workflow clearly:
- raw spectra → metabolites → clinical data → predictive model
⚙️ Machine learning in metabolomics
Three main paradigms:
🔍 Unsupervised learning
- clustering patients/metabolites
- dimensionality reduction (PCA, etc.)
🎯 Supervised learning
- classification (disease vs control)
- regression (risk prediction)
⏳ Specialized models
- survival analysis
- time-to-event predictions
👉 ML is already deeply embedded, even in metabolite identification pipelines.
⚠️ Core challenge: the data itself
Metabolomics is… messy.
1. Curse of dimensionality
- thousands of metabolites
- few samples (“p ≫ n”)
- risk of overfitting
2. Noise & artifacts
- MS produces thousands of signals
- many are:
- background noise
- adducts / fragments
- misannotations
- background noise
👉 can completely distort ML models
3. Missing values
- technical + biological causes
- require complex imputation strategies
🧬 Feature selection & engineering
To survive this complexity, models rely on:
✂️ Feature selection
- filter (statistics)
- wrapper (model-based)
- embedded (e.g., LASSO, RF)
🔄 Feature engineering
- normalization
- scaling
- pathway-based aggregation
👉 This step is absolutely critical for model performance
📊 Clinical evaluation ≠ ML accuracy
This is one of the most important points.
👉 High accuracy ≠ clinical usefulness
Instead, models must optimize:
- sensitivity / specificity
- false positives vs false negatives
- clinical utility metrics (NB, NNB)
👉 because wrong predictions have real consequences
🧠 Explainability problem
Many models are:
❌ black boxes
This is unacceptable in medicine.
👉 Enter XAI (Explainable AI)
- helps understand decisions
- validates biological plausibility
- builds trust
🔗 Pathway analysis as validation
A really nice idea in this paper:
👉 use pathway analysis as an independent check
- confirms biological relevance
- links metabolites → mechanisms
Example:
- Parkinson’s biomarkers
- validated via pathway links to α-synuclein aggregation
🚨 Reality check: current limitations
Despite hype, major issues remain:
- ❌ lack of external validation
- ❌ small, biased datasets
- ❌ poor reproducibility
- ❌ limited interpretability
👉 many models are not clinically ready yet
🧠 Deeper problem: causality
ML finds patterns but:
👉 correlation ≠ causation
To personalize treatment, we need:
- causal inference
- mechanistic understanding
- integration with biology
🚀 Why this matters
This review makes one thing clear:
👉 metabolomics + ML is powerful
👉 but not plug-and-play
Future progress depends on:
- better data quality
- standardized pipelines
- integration with clinical data
- rigorous validation
💚 BioGenies perspective
This fits perfectly with what we care about:
- data quality 🧪
- model interpretability 🧠
- biological grounding 🔬
👉 because good models need good biology—not just good ML