PlastoGram: decoding protein localization inside plastids 🌿🧬
PlastoGram, protein localization, plastids, chloroplast, machine learning, bioinformatics, sequence analysis
📌 Project highlights
- 🌿 Predicts subplastid localization (E, S, TM, TL)
- 🧬 Distinguishes nuclear- vs plastid-encoded proteins
- ⚙️ Ensemble model combining multiple ML approaches
- 🔬 Includes import pathway prediction (Sec / Tat)
- 🚀 Available as web server + R package
🎉 New tool & paper!
👉 understanding where proteins go inside plastids turns out, it’s not trivial 😄
👉 Prediction of protein subplastid localization and origin with PlastoGram
🔗 Try it yourself
👉 plug your sequences in and see where they end up 🌿
🎧 Audio summary
Protein localization inside plastids, multiple compartments, import pathways…
yeah, this gets complicated quickly 😄
👉 Here’s a short audio overview 🎧 explaining what PlastoGram actually does:
🔬 What is this about?
Plastids (like chloroplasts) are:
- 🌿 essential for photosynthesis
- 🧪 central to metabolism
- 🧬 full of proteins from two genomes
👉 nuclear + plastid
But here’s the catch:
👉 proteins don’t just go into plastids, they go to specific sub-compartments
- envelope (E)
- stroma (S)
- thylakoid membrane (TM)
- thylakoid lumen (TL)
And 👉 location = function
⚙️ The core problem
Predicting subplastid localization is hard because:
- 📉 small datasets
- 🔁 high sequence similarity (homology)
- 🧩 overlapping features between classes
👉 especially:
- stromal vs membrane proteins
- nuclear-encoded vs plastid-encoded
🧠 What we built
👉 PlastoGram = ensemble ML model for plastid protein annotation
Key idea:
- break the problem into smaller decisions
- combine them into a final prediction
⚙️ How it works
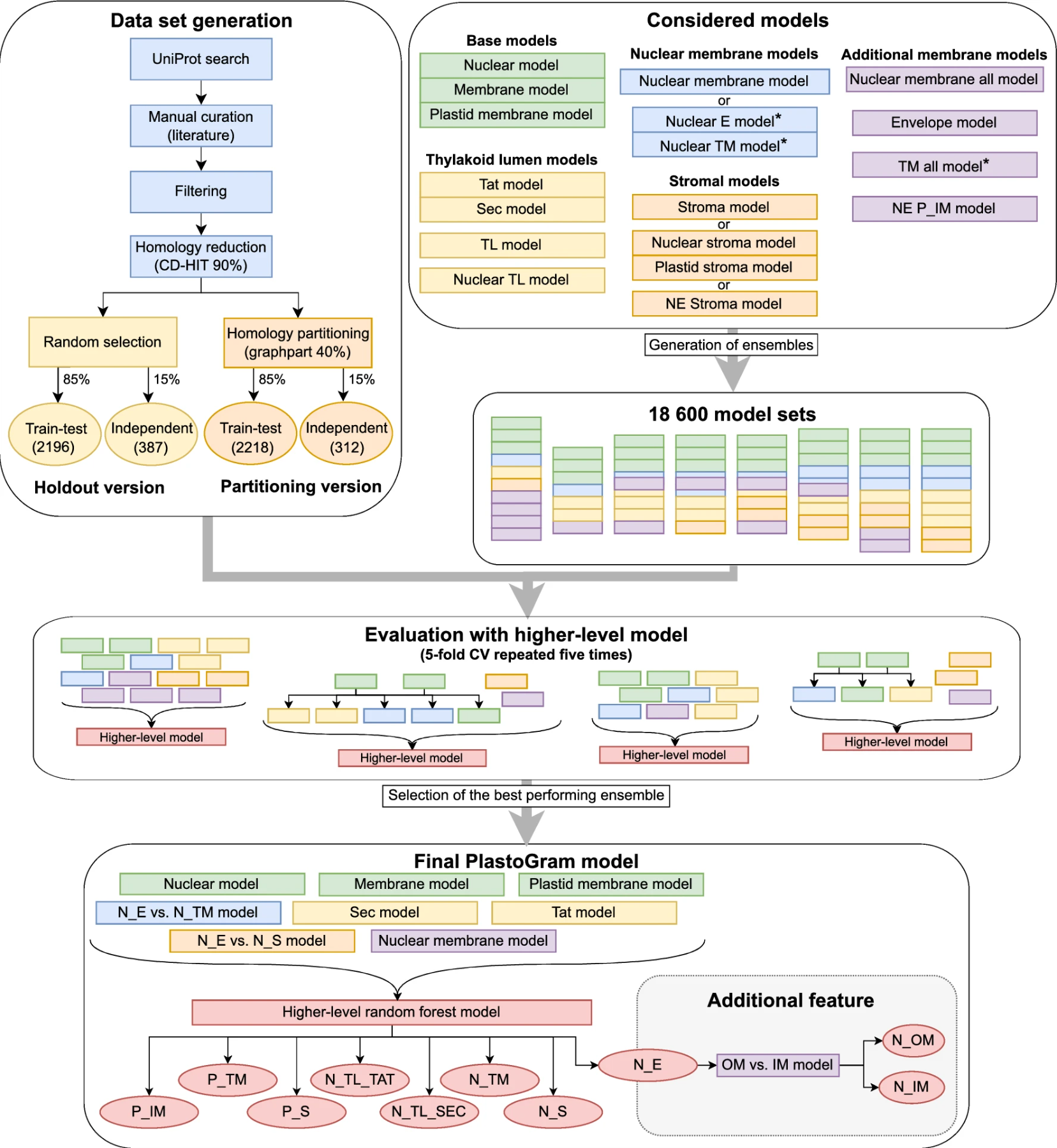
🧩 Ensemble architecture
PlastoGram combines:
- multiple random forest models
- HMM-based models
- a higher-level classifier
👉 stacked together into one system
🔍 What it predicts
For each protein:
- 🧬 origin:
- nuclear
- plastid
- nuclear
- 🌿 localization:
- envelope
- stroma
- thylakoid membrane
- thylakoid lumen
- envelope
- 🚪 (if TL):
- Sec pathway
- Tat pathway
- Sec pathway
🧪 Data matters (a lot)
You built:
- manually curated dataset
- thousands of proteins
- careful filtering & homology control
👉 because garbage in = garbage out
📊 Key insights from the paper
⚠️ Data is still the bottleneck
- some classes have <50 proteins
- limits model reliability
👉 especially for rare compartments
🧬 Plastid vs nuclear proteins behave differently
- plastid-encoded → easier to predict
- nuclear-encoded → more complex
👉 due to targeting signals & diversity
🤯 Some classes are inherently hard
Example:
- outer membrane vs stroma
👉 almost indistinguishable in features
👉 even PCA shows strong overlap
🧠 Models learn real biology
Nice example:
- n-grams capture known motifs (e.g. targeting signals)
👉 ML is not just guessing, it learns biology
🏆 Performance
- strong improvement over baseline models
- competitive vs existing tools (e.g. SChloro)
- especially good for:
- plastid-encoded proteins
- abundant classes
- plastid-encoded proteins
🚀 Why this matters
Protein localization is:
👉 fundamental for:
- functional annotation
- pathway reconstruction
- synthetic biology
And PlastoGram enables:
👉 more precise, automated annotation
💚 BioGenies perspective
This project is a perfect example of:
- 🧠 combining biology + ML
- ⚙️ building usable tools (not just models)
- 🔬 caring about data quality
👉 because better annotations → better biology