Amyloid databases: mapping the aggregation universe 🧬📊
amyloids, protein aggregation, databases, bioinformatics, amyloid prediction, protein misfolding, aggregation
📌 Project highlights
- 🧬 Comprehensive overview of amyloid & aggregation databases
- 📊 Covers sequence, structural and interaction resources
- 🔗 Highlights connections between databases and prediction tools
- ⚠️ Identifies key limitations in current resources
- 🚀 Provides curated list of databases: link
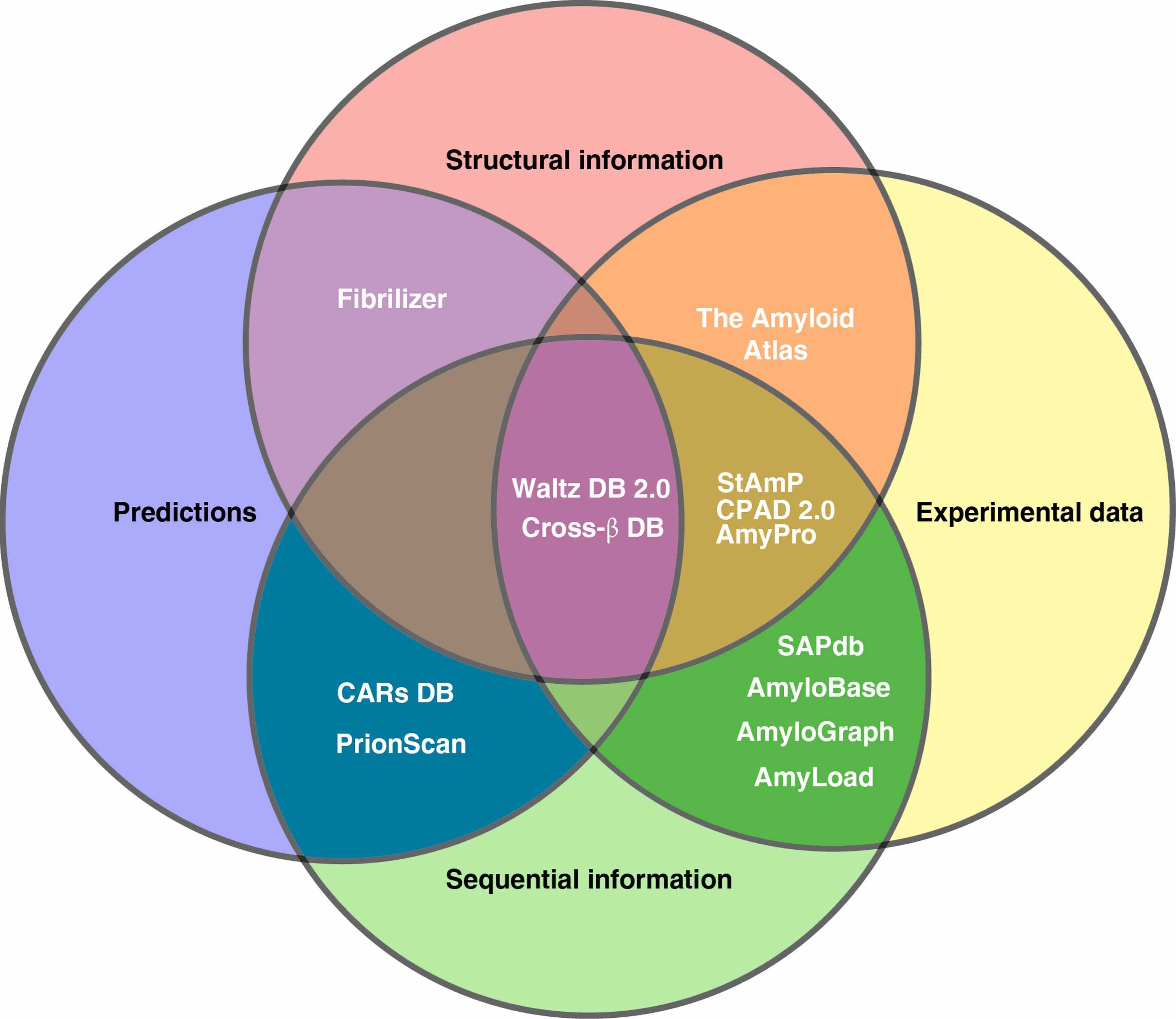
🎉 New review out! This one is less about a single tool and more about the entire ecosystem of amyloid data 😄
🔗 Explore the resources
👉 This is basically a map of the amyloid bioinformatics landscape.
🎧 Audio summary
Too many databases to remember? Same 😄
👉 We’ve added a short audio overview 🎧 so you don’t have to memorize all of them.
🔬 What is this about?
Amyloid aggregation is a complex, multi-factorial process involving:
- sequence features
- 3D structure
- environmental conditions
👉 and it underlies:
- neurodegenerative diseases
- biotechnological challenges
- functional biological processes
Because of this complexity:
👉 researchers have built many specialized databases to organize experimental knowledge
🧠 What we reviewed
We systematically analyzed amyloid-related databases, grouping them into:
🧬 Sequence-based databases
- focus on aggregation-prone regions (APRs)
- example: AmyLoad, AmyPro
🧊 Structure-based databases
- store 3D fibril structures
- example: Amyloid Atlas
🔗 Interaction databases
- capture cross-interactions between amyloids
- example: AmyloGraph
👉 Each database captures different aspects of aggregation.
📊 Key insight: fragmentation problem
There is no single “perfect” database.
Instead:
- each resource focuses on a specific niche
- data formats and annotations differ
- integration is difficult
👉 Result:
❌ no unified benchmark dataset
❌ hard to compare prediction tools
❌ fragmented knowledge
⚙️ Databases ↔︎ prediction tools (the feedback loop)
One of the most important conclusions:
👉 databases and prediction tools co-evolve
- experimental datasets → enable model development
- prediction tools → generate new hypotheses
- new experiments → expand databases
👉 A continuous feedback loop driving the field forward.
🧬 Examples of this interplay
- AmyloGraph → enabled PACT / AmyloComp (cross-interactions)
- AmyloBase → contributed to AGGRESCAN
- Waltz datasets → led to WALTZ algorithm
👉 Data → model → better data → better model
⚠️ Key limitations (important!)
Across databases:
- 🔍 limited search & filtering
- 📤 poor export options
- 🧾 incomplete metadata
- 🤖 reliance on predictions (with biases)
👉 And most importantly: aggregation is not only sequence-dependent
Environmental factors matter:
- pH
- temperature
- concentration
- cofactors
🚀 Why this matters
This review shows:
👉 we have a lot of data
👉 but not yet fully integrated knowledge
Future directions:
- better standardization (e.g. MIRRAGGE)
- integration of datasets
- ML models using multi-dimensional data
💚 BioGenies perspective
This is basically our playground:
- AmyloGraph 🔗
- CARs DB 🧬
- aggregation prediction 🧠
👉 And a key takeaway we strongly agree with data integration is the next big step in amyloid research
